Creating synthetic antibodies to detect molecules
Synthetic polymers coating a nanoparticle surface can recognize
specific molecules, just like an antibody, for detecting
neurotransmitters, diseases, or environmental toxins, for example
November 27, 2013
[+]
MIT chemical
engineers have developed a novel way to generate nanoparticles that can
recognize specific molecules, opening up a new approach to building
durable sensors for many different compounds, among other applications.
MIT
chemical engineers created this sensor that can recognize riboflavin by
coating a carbon nanotube with amphiphilic polymers (credit: Jingqing
Zhang et al.)
To create these “synthetic antibodies,” the researchers used carbon nanotubes — hollow, nanometer-thick cylinders made of carbon that naturally fluoresce when exposed to laser light.
The MIT team coated the nanotubes with specifically designed amphiphilic polymers — polymers that are drawn to both oil and water, like soap. This approach offers a huge array of recognition sites specific to different targets, and could be used to create sensors to monitor diseases such as cancer, inflammation, or diabetes in living systems.
“This new technique gives us an unprecedented ability to recognize any target molecule by screening nanotube-polymer complexes to create synthetic analogs to antibody function,” says Michael Strano, the Carbon P. Dubbs Professor of Chemical Engineering at MIT and senior author of the study, which appears in the Nov. 24 online edition of Nature Nanotechnology.
Sensors targeted to specific molecules
This approach can provide a more durable alternative to coating sensors such as carbon nanotubes with actual antibodies, which can break down inside living cells and tissues. Another family of commonly used recognition molecules are DNA aptamers, which are short pieces of DNA that interact with specific targets, depending on the aptamer sequence. However, there are not aptamers specific to many of molecules that one might want to detect, Strano says.
In the new paper, the researchers describe molecular recognition sites that enable the creation of sensors specific to riboflavin, estradiol (a form of estrogen), and L-thyroxine (a thyroid hormone), but they are now working on sites for many other types of molecules, including neurotransmitters, carbohydrates, and proteins.
[+]
Their approach takes advantage of a phenomenon that occurs when
certain types of polymers bind to a carbon nanotube. These polymers,
known as amphiphilic, have both hydrophobic (water-rejecting) and
hydrophilic (water-accepting) regions.
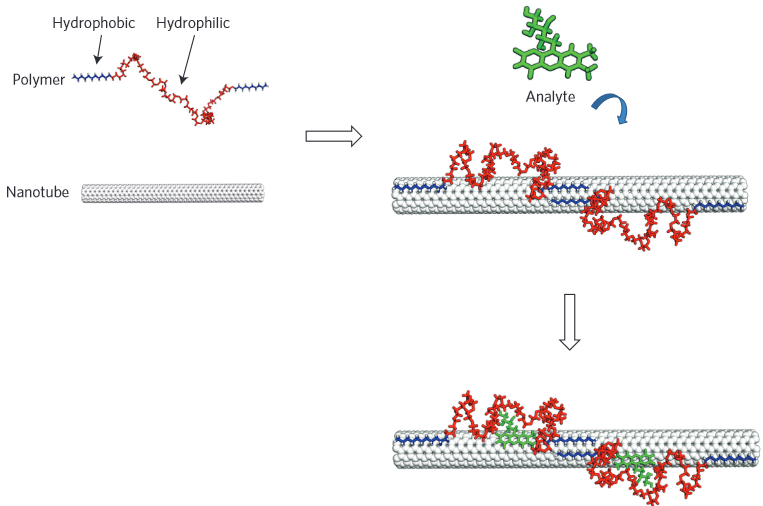
Schematic
of the molecular recognition concept. A polymer with an alternating
hydrophobic and hydrophilic sequence adopts a specific conformation when
adsorbed to the nanotube. The polymer is pinned in place to create a
selective molecular recognition site for the analyte (molecule of
interest), leading to either a wavelength or intensity change in
single-wall nanotube fluorescence. (Credit: Jingqing Zhang et al./Nature Nanotechnology)
These polymers are designed and synthesized such that when the polymers are exposed to carbon nanotubes, the hydrophobic regions latch onto the tubes like anchors and the hydrophilic regions form a series of loops extending away from the tubes.
These loops form a new layer surrounding the nanotube, known as a corona. The MIT researchers found that the loops within the corona are arranged very precisely along the tube, and the spacing between the anchors determines which target molecule will be able to wedge into the loops and alter the carbon nanotube’s fluorescence.
“We hope that our sensors will provide the scientific community with a new approach to detecting important molecules that is rapid and label-free,” said Markita del Carpio Landry, Ph.D., lead author, in an email interview with KurzweilAI.
“The tools required to make these sensors are relatively inexpensive and accessible, so we hope other research groups can use this technology to support their day-to-day research.
“For instance, if we can design sensors to detect molecules of clinical importance, they could be used to design technologies to sense bio-molecular markers that are indicative of diseases. Alternatively, we could design sensors to recognize hazardous molecules in our environments, such as environmental pollutants.
“Since the ability to design a sensor is only limited by our imaginations (instead of by available naturally occurring antibodies), the potential applications are equally numerous!”
The research was funded by the National Science Foundation and the Army Research Office through MIT’s Institute for Soldier Nanotechnologies.
Abstract of Nature Nanotechnology paper
Understanding molecular recognition is of fundamental importance in applications such as therapeutics, chemical catalysis and sensor design. The most common recognition motifs involve biological macromolecules such as antibodies and aptamers. The key to biorecognition consists of a unique three-dimensional structure formed by a folded and constrained bioheteropolymer that creates a binding pocket, or an interface, able to recognize a specific molecule. Here, we show that synthetic heteropolymers, once constrained onto a single-walled carbon nanotube by chemical adsorption, also form a new corona phase that exhibits highly selective recognition for specific molecules. To prove the generality of this phenomenon, we report three examples of heteropolymer-nanotube recognition complexes for riboflavin, L-thyroxine and oestradiol. In each case, the recognition was predicted using a two-dimensional thermodynamic model of surface interactions in which the dissociation constants can be tuned by perturbing the chemical structure of the heteropolymer. Moreover, these complexes can be used as new types of spatiotemporal sensors based on modulation of the carbon nanotube photoemission in the near-infrared, as we show by tracking riboflavin diffusion in murine macrophages.
(¯`*• Global Source and/or more resources at http://goo.gl/zvSV7 │ www.Future-Observatory.blogspot.com and on LinkeIn Group's "Becoming Aware of the Futures" at http://goo.gl/8qKBbK │ @SciCzar │ Point of Contact: www.linkedin.com/in/AndresAgostini
 Washington
Washington